Loading required package: Matrix
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, union… and why random intercepts are often not enough
Daniel S. Mazhari-Jensen
July 23, 2025

In this post, we’ll discuss why random slopes are often of special concern in health science data and biometrics. We will use lme4 and ggplot2 to do so. The data set we’ll use to demonstrate this comes from …
Loading required package: Matrix
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionset.seed(314)
# Parameters
n_subjects <- 30
n_sessions <- 5
# Fixed effect of session: general learning effect
beta_0 <- 600 # Average baseline RT in ms
beta_1 <- -10 # Average decrease in RT per session
# Random effects: standard deviations
sd_intercept <- 50 # SD of baseline RTs across subjects
sd_slope <- 25 # SD of learning rates (slopes)
rho <- 0.2 # Correlation between intercepts and slopes
# Within-subject residual error
sigma_error <- 30
# Construct subject-level random effects (correlated)
subject_re <- MASS::mvrnorm(
n = n_subjects,
mu = c(0, 0),
Sigma = matrix(c(sd_intercept^2, rho*sd_intercept*sd_slope,
rho*sd_intercept*sd_slope, sd_slope^2), 2)
) |>
as.data.frame() |>
setNames(c("intercept_re", "slope_re")) |>
dplyr::mutate(subject = factor(1:n_subjects))
# Create session vector repeated for each subject
session <- rep(0:(n_sessions - 1), times = n_subjects)
# Repeat subject IDs for each session
subject <- rep(subject_re$subject, each = n_sessions)
# Join the random effects
sim_data <- tibble::tibble(subject = subject, session = session) |>
dplyr::left_join(subject_re, by = "subject") |>
dplyr::mutate(
reaction_time = beta_0 + intercept_re +
(beta_1 + slope_re) * session +
rnorm(dplyr::n(), mean = 0, sd = sigma_error)
)
# View a few rows
head(sim_data)# A tibble: 6 × 5
subject session intercept_re slope_re reaction_time
<fct> <int> <dbl> <dbl> <dbl>
1 1 0 63.5 14.4 663.
2 1 1 63.5 14.4 703.
3 1 2 63.5 14.4 673.
4 1 3 63.5 14.4 691.
5 1 4 63.5 14.4 660.
6 2 0 -39.0 15.7 566.tibble [150 × 5] (S3: tbl_df/tbl/data.frame)
$ subject : Factor w/ 30 levels "1","2","3","4",..: 1 1 1 1 1 2 2 2 2 2 ...
$ session : int [1:150] 0 1 2 3 4 0 1 2 3 4 ...
$ intercept_re : num [1:150] 63.5 63.5 63.5 63.5 63.5 ...
$ slope_re : num [1:150] 14.4 14.4 14.4 14.4 14.4 ...
$ reaction_time: num [1:150] 663 703 673 691 660 ...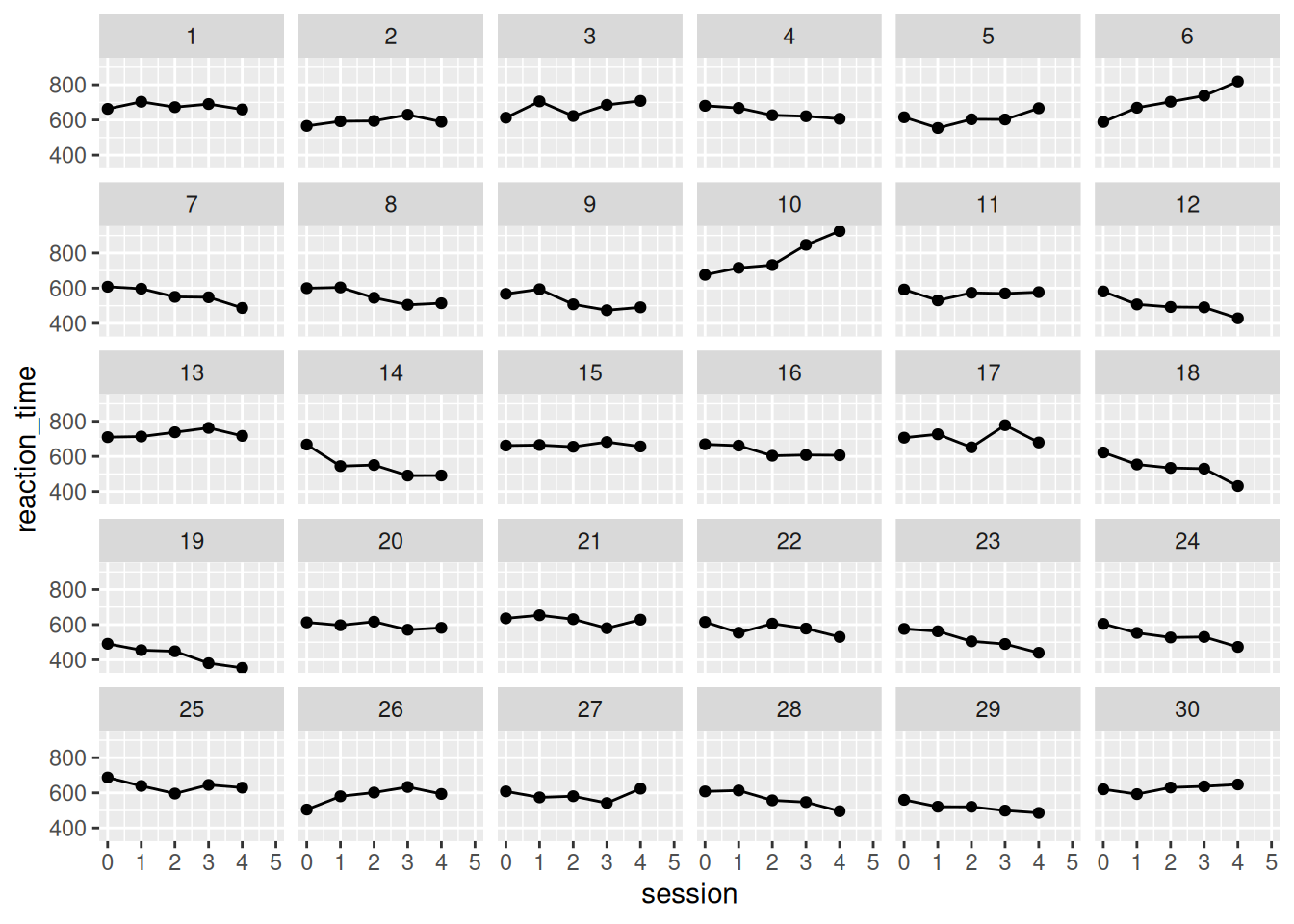
Linear mixed model fit by REML ['lmerMod']
Formula: reaction_time ~ 1 + session + (1 | subject)
Data: sim_data
REML criterion at convergence: 1639.3
Scaled residuals:
Min 1Q Median 3Q Max
-2.6679 -0.4728 -0.0547 0.4959 3.8078
Random effects:
Groups Name Variance Std.Dev.
subject (Intercept) 5402 73.50
Residual 2103 45.85
Number of obs: 150, groups: subject, 30
Fixed effects:
Estimate Std. Error t value
(Intercept) 614.455 14.904 41.229
session -7.529 2.647 -2.844
Correlation of Fixed Effects:
(Intr)
session -0.355[1] 0.08115909Linear mixed model fit by REML ['lmerMod']
Formula: reaction_time ~ 1 + session + (session | subject)
Data: sim_data
REML criterion at convergence: 1550.6
Scaled residuals:
Min 1Q Median 3Q Max
-2.06073 -0.52670 -0.00823 0.53737 2.59156
Random effects:
Groups Name Variance Std.Dev. Corr
subject (Intercept) 2090.4 45.72
session 565.9 23.79 0.30
Residual 723.6 26.90
Number of obs: 150, groups: subject, 30
Fixed effects:
Estimate Std. Error t value
(Intercept) 614.455 9.174 66.981
session -7.529 4.612 -1.632
Correlation of Fixed Effects:
(Intr)
session 0.147 [1] 0.1532919
Call:
lm(formula = Reaction ~ 1 + Days, data = sleepstudy)
Residuals:
Min 1Q Median 3Q Max
-110.848 -27.483 1.546 26.142 139.953
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 251.405 6.610 38.033 < 2e-16 ***
Days 10.467 1.238 8.454 9.89e-15 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 47.71 on 178 degrees of freedom
Multiple R-squared: 0.2865, Adjusted R-squared: 0.2825
F-statistic: 71.46 on 1 and 178 DF, p-value: 9.894e-15Here is the model prediction - notice how every participant is assumed to have the same initial value and linear trajectory. This is due to pooling, an assumption from ANOVA.
If we want to be more sensitive to individual variability, we need to allow participants to be modeled more individually. This can be done by computing linear regression for each participant, treating them as individual cases with no overlap.
Call:
Model: Reaction ~ Days | NULL
Data: sleepstudy
Coefficients:
(Intercept)
Estimate Std. Error t value Pr(>|t|)
308 244.1927 15.04169 16.23439 2.419368e-34
309 205.0549 15.04169 13.63244 1.067180e-27
310 203.4842 15.04169 13.52802 1.993900e-27
330 289.6851 15.04169 19.25882 1.122068e-41
331 285.7390 15.04169 18.99647 4.646933e-41
332 264.2516 15.04169 17.56795 1.236403e-37
333 275.0191 15.04169 18.28379 2.303436e-39
334 240.1629 15.04169 15.96649 1.135574e-33
335 263.0347 15.04169 17.48705 1.946826e-37
337 290.1041 15.04169 19.28667 9.653936e-42
349 215.1118 15.04169 14.30104 1.983389e-29
350 225.8346 15.04169 15.01391 2.939145e-31
351 261.1470 15.04169 17.36155 3.943049e-37
352 276.3721 15.04169 18.37374 1.402577e-39
369 254.9681 15.04169 16.95077 4.023936e-36
370 210.4491 15.04169 13.99106 1.253782e-28
371 253.6360 15.04169 16.86221 6.656453e-36
372 267.0448 15.04169 17.75365 4.373979e-38
Days
Estimate Std. Error t value Pr(>|t|)
308 21.764702 2.817566 7.7246464 1.741840e-12
309 2.261785 2.817566 0.8027444 4.234454e-01
310 6.114899 2.817566 2.1702769 3.162541e-02
330 3.008073 2.817566 1.0676139 2.874813e-01
331 5.266019 2.817566 1.8689956 6.365457e-02
332 9.566768 2.817566 3.3954013 8.857738e-04
333 9.142045 2.817566 3.2446604 1.462120e-03
334 12.253141 2.817566 4.3488388 2.574673e-05
335 -2.881034 2.817566 -1.0225257 3.082469e-01
337 19.025974 2.817566 6.7526272 3.315759e-10
349 13.493933 2.817566 4.7892159 4.115160e-06
350 19.504017 2.817566 6.9222924 1.356856e-10
351 6.433498 2.817566 2.2833528 2.387301e-02
352 13.566549 2.817566 4.8149886 3.683105e-06
369 11.348109 2.817566 4.0276282 9.081880e-05
370 18.056151 2.817566 6.4084212 1.964766e-09
371 9.188445 2.817566 3.2611283 1.385338e-03
372 11.298073 2.817566 4.0098697 9.718197e-05
Residual standard error: 25.59182 on 144 degrees of freedomWe can relax the assumption of pooling by allowing each participant to have their own intercept.
Linear mixed model fit by REML ['lmerMod']
Formula: Reaction ~ 1 + Days + (1 | Subject)
Data: sleepstudy
REML criterion at convergence: 1786.5
Scaled residuals:
Min 1Q Median 3Q Max
-3.2257 -0.5529 0.0109 0.5188 4.2506
Random effects:
Groups Name Variance Std.Dev.
Subject (Intercept) 1378.2 37.12
Residual 960.5 30.99
Number of obs: 180, groups: Subject, 18
Fixed effects:
Estimate Std. Error t value
(Intercept) 251.4051 9.7467 25.79
Days 10.4673 0.8042 13.02
Correlation of Fixed Effects:
(Intr)
Days -0.371Note the fixed effect 10.4673 / 0.8042 = 13.0157921
Linear mixed model fit by REML ['lmerMod']
Formula: Reaction ~ 1 + Days + (1 + Days | Subject)
Data: sleepstudy
REML criterion at convergence: 1743.6
Scaled residuals:
Min 1Q Median 3Q Max
-3.9536 -0.4634 0.0231 0.4634 5.1793
Random effects:
Groups Name Variance Std.Dev. Corr
Subject (Intercept) 612.10 24.741
Days 35.07 5.922 0.07
Residual 654.94 25.592
Number of obs: 180, groups: Subject, 18
Fixed effects:
Estimate Std. Error t value
(Intercept) 251.405 6.825 36.838
Days 10.467 1.546 6.771
Correlation of Fixed Effects:
(Intr)
Days -0.138Note the fixed effect 10.467 / 1.546 = 6.7703752
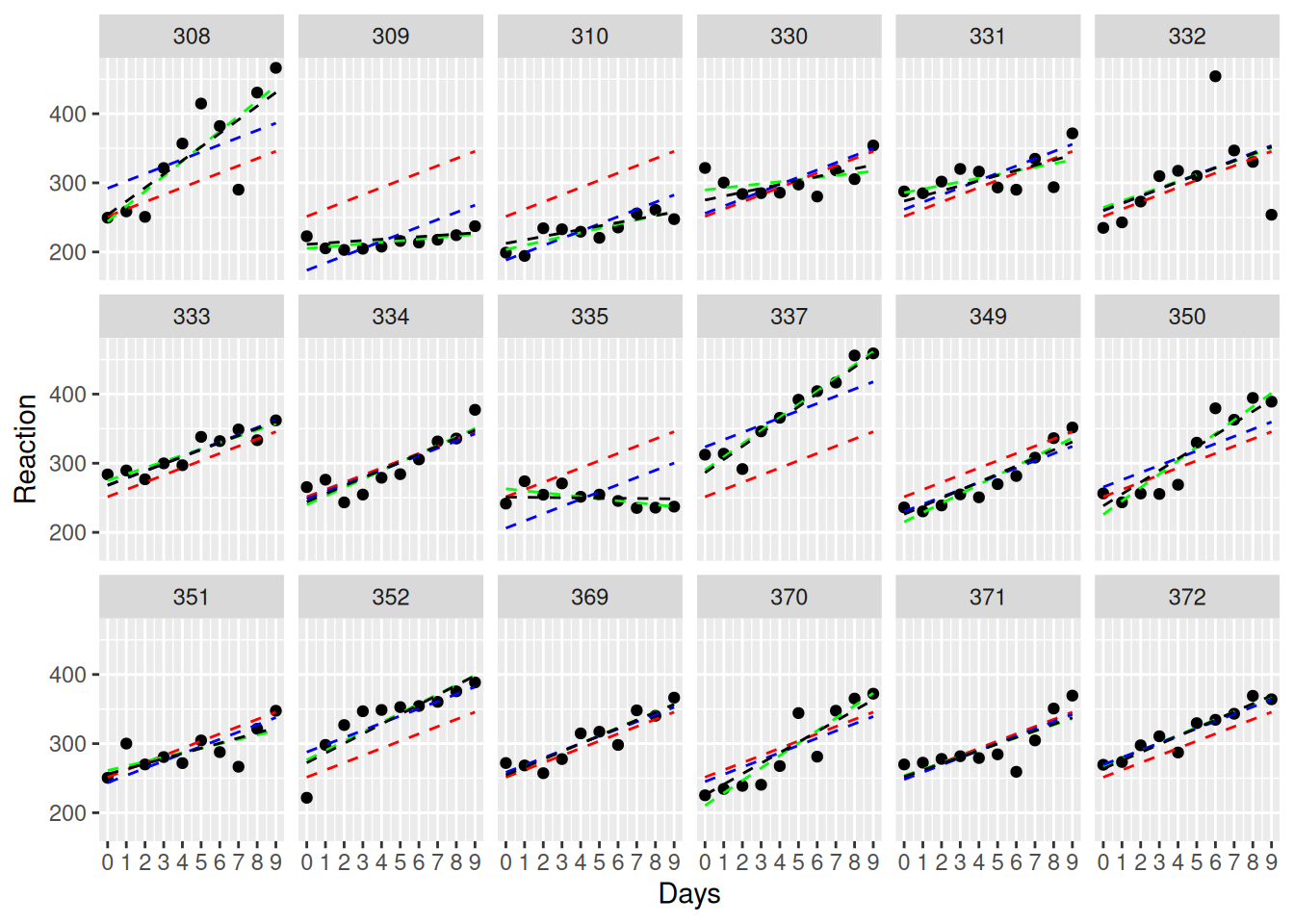
ggplot(sleepstudy, aes(Days, Reaction, group=Subject)) +
facet_wrap(~Subject, ncol=6) +
geom_point() +
geom_line(aes(y=fitted(anova_approach)), linetype=2, color = "red") +
geom_line(aes(y=fitted(no_pooling)), linetype=2, color = "green") +
geom_line(aes(y=fitted(RI_only)), linetype=2, color = "blue") +
geom_line(aes(y=fitted(RI_RS_corr)), linetype=2, color = "black") +
scale_x_continuous(limits=c(0, 9),breaks=c(0:9))